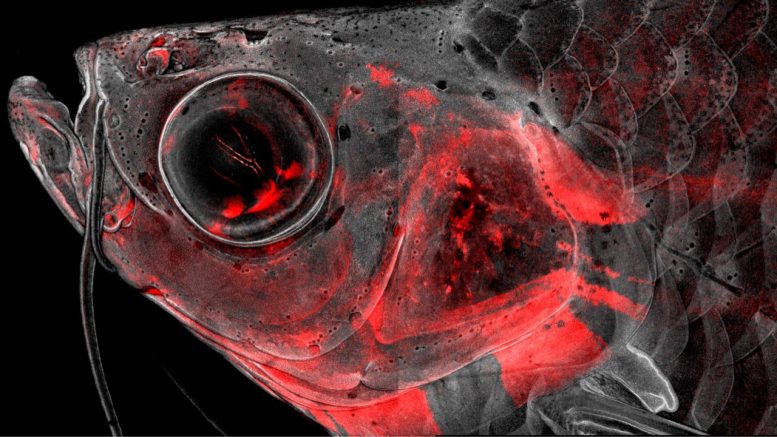
Confocal microscopy image of an adult zebrafish head with neural crest-derived cells in red. The Crump laboratory has actually used single-cell sequencing to comprehend how these cells repair the head and construct skeleton, with ramifications for understanding human craniofacial birth flaws and enhancing repair of skeletal tissues. Credit: Image by Hun-Jhen Chen/Crump Lab
Cranial neural crest cells, or CNCCs, contribute to numerous more body parts than their humble name suggests. These impressive stem cells not just form most of the skull and facial skeleton in all vertebrates ranging from fish to human beings, however likewise can create everything from gills to the cornea.
” CNCCs have actually long fascinated biologists by the extraordinary diversity of cell types they can produce. By studying this procedure in the genetically tractable zebrafish, we have actually recognized much of the possible switches that permit CNCCs to form these really various cell types,” stated Gage Crump, teacher of stem cell biology and regenerative medication at the Keck School of Medicine of USC.
Led by postdoc Peter Fabian and PhD students Kuo-Chang Tseng, Mathi Thiruppathy, and Claire Arata, the group of researchers permanently labeled CNCCs with a red fluorescent protein to track which cell types came from CNCCs throughout the lifetime of zebrafish. They then utilized an effective kind of technique, known as “single-cell genomics,” to identify the complete set of active genes and the organization of the DNA across hundreds of thousands of specific CNCCs. The enormous amount of information generated required the scientists to establish a new computational tool to understand it.
The Crump laboratory has used single-cell sequencing to understand how these cells develop and repair the head skeleton, with ramifications for comprehending human craniofacial birth problems and enhancing repair of skeletal tissues. Cranial neural crest cells, or CNCCs, contribute to lots of more body parts than their modest name suggests. Led by postdoc Peter Fabian and PhD trainees Kuo-Chang Tseng, Mathi Thiruppathy, and Claire Arata, the team of scientists permanently identified CNCCs with a red fluorescent protein to keep track of which cell types came from CNCCs throughout the lifetime of zebrafish. Through this new bioinformatic technique, the team discovered that CNCCs do not begin out with all the details required to make the huge diversity of cell types.
” We developed a kind of computational analysis that we called Constellations, because the final visual output of the strategy is reminiscent of constellations of stars in the sky,” stated Fabian. “In contrast to astrology, our Constellations algorithm actually can forecast the future of cells and expose the crucial genes that likely control their advancement.”
Through this brand-new bioinformatic approach, the group discovered that CNCCs do not begin out with all the details required to make the substantial variety of cell types. Real-life experiments verified that Constellations properly pinpointed the function of a family of “FOX” genes in facial cartilage formation, and a previously unappreciated function for “GATA” genes in the development of gill breathing cell types that permit fish to breathe.
” By carrying out one of the most comprehensive single-cell research studies of a vertebrate cell population to date, we not just gained considerable insights into the advancement of the vertebrate head, however likewise created a broadly useful computational tool for studying the development and regrowth of organ systems throughout the body,” said Crump.
Reference: “Lifelong single-cell profiling of cranial neural crest diversification in zebrafish” by Peter Fabian, Kuo-Chang Tseng, Mathi Thiruppathy, Claire Arata, Hung-Jhen Chen, Joanna Smeeton, Nellie Nelson and J. Gage Crump, 10 January 2022, Nature Communications.DOI: 10.1038/ s41467-021-27594-w.
Extra co-authors in the Crump Lab included PhD trainee Hung-Jhen Chen, postdoc Joanna Smeeton, and research specialist Nellie Nelson. Smeeton is now an assistant professor at Columbia University, and Nelson is a PhD student at the University of California, Irvine.
The research was federally moneyed by the National Institutes of Health (grants NIDCR R35 DE027550, NIDCR K99 DE029858, NIDCR F31 DE029682-02, NICHD T32 HD060549).