The researchers report their findings in the journal Science.
Matthew McCallum, a postdoctoral fellow in Veeslers lab, and Nadine Czudnochowski, a Vir Biotechnology scientist, were lead authors on the paper.
The omicron variation, which was initially recognized in November 2021 in South Africa, is causing a surge of infections worldwide. In addition to being highly transmittable, the version can evade antibodies versus earlier variations causing advancement infections amongst those who have been vaccinated and those who have actually been contaminated formerly.
The infectiousness of the virus is believed to be at least in part due to the big number of mutations in the amino acid sequences of the viruss spike protein. The infection uses the spike protein to latch on to and go into the cells it contaminates. The omicron spike protein has 37 anomalies that identify it from the first SARS-CoV-2 isolates in 2020.
Previous research by Veesler and associates have actually shown that antibodies created by the six most commonly utilized vaccines, and all but one of monoclonal antibodies presently utilized to deal with infections, have actually a lowered or abrogated ability to neutralize omicron.
However a number of the anomalies in the variant affect the structure of the region of the spike protein that is responsible for connecting to and entering cells, an area called the receptor binding domain, and lots of anticipated the resulting modifications in the receptor binding domain structure may impair the capability of the variant to bind to its target on cells. This target is protein called angiotensin converting enzyme-2, or ACE2. In their research study, Veesler and his associates found that the changes had actually increased the capability of the receptor binding domain to bind to ACE2 by 2.4-fold.
To understand how omicron accumulated so many anomalies while keeping effective interactions with the host receptor ACE2, Veesler and his coworkers utilized cryo-electron tiny and X-ray crystallographic research studies to unveil the 3D company of the omicron spike protein. The researchers likewise determined how the structural changes in the spike protein affected the capability of antibodies effective against previous variants to bind to Omicron.
Utilizing these techniques, the researchers reveal how the mutations changed how the protein engages with antibodies so that the capability of nearly all monoclonal antibodies against it is decreased, while, at the exact same time the capability of the spike receptor-binding domain to bind ACE2 is enhanced. The general impact has been to make it possible for the receptor binding domain to evade antibodies targeting it and to bind to ACE2 even more firmly.
The findings show what a powerful opponent SARS-CoV-2 is, states Veesler.
” This infection has extraordinary plasticity: It can alter a lot and still keep all the functions it requires to infect and duplicate,” he noted. “And its almost guaranteed omicron is not the last version were going to see.”
The objective going forward must be to focus on and determine additional regions on the spike protein that can not be changed without triggering the protein to lose function, Veesler said. These areas tend to stay saved even as other parts of the protein mutates since of their importance.
Such conserved areas of viral proteins are therefore most likely to stay unchanged in any brand-new version that might emerge. These areas would make perfect targets for new vaccines and rehabs that might be effective not only versus new versions but new sarbecoviruses, the group of viruses consisting of SARS-CoV-2 and SARS-CoV, Veesler said.
Recommendation: “Structural basis of SARS-CoV-2 omicron immune evasion and receptor engagement” 25 January 2022, Science.DOI: 10.1126/ science.abn8652.
The research study was supported by the National Institute of Health, the National Institute of Allergy and Infectious Disease, the National Institute of General Medical Sciences, the Burroughs Wellcome Fund, Fast Grants, the University of Washington Arnold and Mabel Beckman cryoEM center, the Howard Hughes Medical Institute, the Wellcome Trust and a Pew Biomedical Scholars Award.
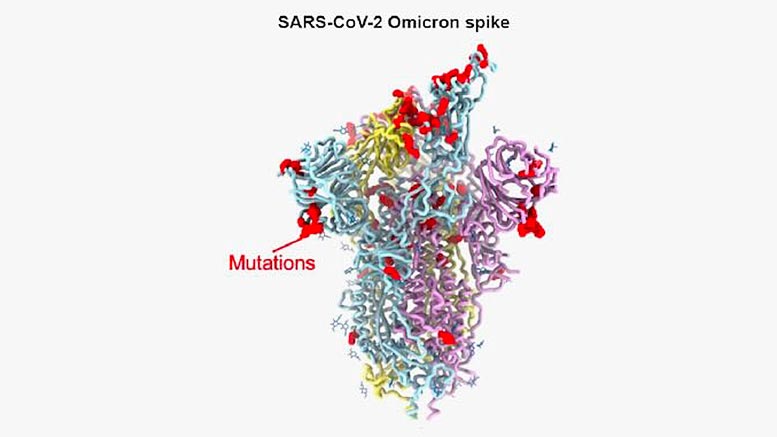
Design of the omicron alternative spike protein reveals the place of a few of its 37 mutations (red spheres). Credit: David Veesler Lab
Findings describe how mutations in the protein permit the omicron variation of the pandemic coronavirus to evade antibodies against previous versions yet stay so infectious.
A global group of researchers has determined the exact structural modifications in the spike protein of the omicron variation. Their observations discuss how the infection is able to evade antibodies against previous versions and still remain extremely transmittable.
” The findings provide a blueprint that researchers can use to develop brand-new countermeasures, whether they be therapies or vaccines, against omicron and other coronavirus variations that may emerge,” stated David Veesler, investigator with the Howard Hughes Medical Institute and associate teacher of biochemistry at the University of Washington School of Medicine in Seattle. He led the research effort with Gyorgy Snell from Vir Biotechnology, Inc. in San Francisco.
The infectiousness of the infection is thought to be at least in part due to the big number of anomalies in the amino acid series of the viruss spike protein. The omicron spike protein has 37 mutations that differentiate it from the first SARS-CoV-2 isolates in 2020.
Numerous of the anomalies in the alternative impact the structure of the region of the spike protein that is responsible for attaching to and getting in cells, a region called the receptor binding domain, and lots of anticipated the resulting modifications in the receptor binding domain structure might impair the capability of the alternative to bind to its target on cells. To understand how omicron collected so lots of anomalies while keeping effective interactions with the host receptor ACE2, Veesler and his coworkers used cryo-electron microscopic and X-ray crystallographic studies to unveil the 3D organization of the omicron spike protein. The scientists also figured out how the structural modifications in the spike protein affected the capability of antibodies reliable versus previous variations to bind to Omicron.